
May 09, 2026
Nanopore sequencing stands as one of the most elegant and disruptive innovations in genomics. Rather than relying on chemical reactions or light to read DNA, it listens to the subtle electrical whispers of individual molecules as they thread through a tiny protein pore. This third-generation technology delivers real-time, long-read data that has transformed everything from plasmid verification to complex genome assembly.
At Quintara Bio, we harness the latest Oxford Nanopore platforms to offer rapid, cost-effective nanopore sequencing services — from same-day plasmid results to high-depth amplicon analysis — so researchers can focus on discovery instead of instrument management.
Nanopore sequencing is a single-molecule, real-time DNA and RNA sequencing method that identifies nucleotides by measuring changes in electrical current as nucleic acids pass through a biological nanopore. Developed by Oxford Nanopore Technologies, it belongs firmly to the third-generation sequencing category, distinct from earlier methods that depend on amplification or synthesis.
At its heart, the technology does something beautifully simple yet profound: it converts molecular events directly into electrical signals. Instead of breaking DNA into short fragments and rebuilding the sequence computationally, nanopore sequencing reads long, native strands as they move through a nanoscale channel. This principle enables ultra-long reads, preserves natural modifications, and provides data the moment sequencing begins.
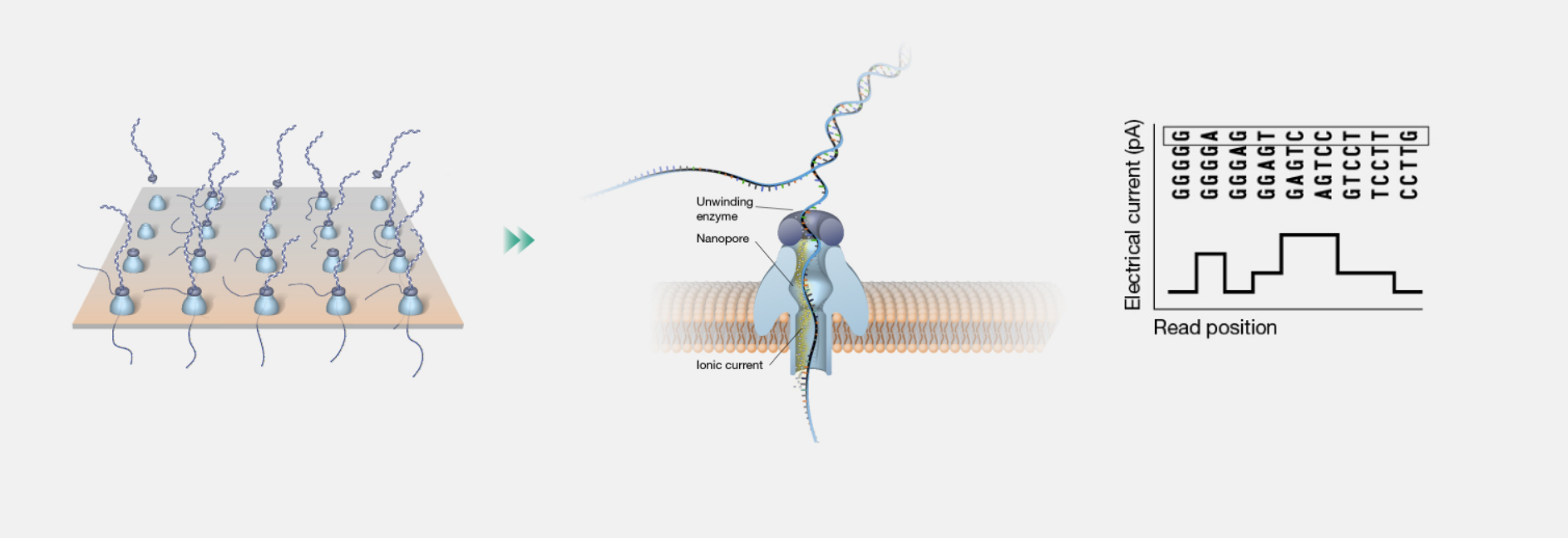
The magic of nanopore sequencing lies in a straightforward biophysical process. DNA and RNA molecules carry a natural negative charge. When an electric field is applied across a membrane containing a nanopore, these molecules are pulled through the pore like thread through the eye of a needle.
A steady ionic current flows through the open nanopore under normal conditions. As each base (or short group of bases known as a k-mer) enters the narrow constriction of the pore, it partially blocks the flow of ions, causing a measurable disruption in the current. Each nucleotide — A, C, G, or T — produces a unique signature because of its size, shape, and charge. These disruptions are captured thousands of times per second as a continuous electrical trace.
Importantly, the sequencer does not "read" DNA directly. It detects the raw electrical signal and relies on sophisticated software to interpret what that signal means. This indirect but highly effective approach is what allows nanopore sequencing to work with native, unmodified molecules.
A nanopore sequencing device is remarkably straightforward yet precisely engineered. The core component is a flow cell containing a synthetic polymer membrane immersed in an ionic solution. Embedded in this membrane are thousands to millions of nanopore proteins — modified transmembrane channel proteins often called "reader" proteins.
Each nanopore sits above its own electrode and sensor on a chip. The membrane acts as a high-resistance barrier, forcing all current to flow exclusively through the nanopores when voltage is applied. This setup creates a sensitive electrical circuit where even tiny changes in ion flow produce detectable signals.
The entire system is compact enough that the MinION device fits in the palm of your hand and connects via USB, while larger PromethION systems scale to population-level throughput. At Quintara Bio, we optimize these flow cells with the latest R10.4.1 chemistry to maximize yield and accuracy for every customer sample.
The workflow is designed for speed and simplicity, making it accessible for both routine and complex projects.
High-quality DNA or RNA is extracted and, depending on the application, may be fragmented or kept intact for ultra-long reads. Adapters containing motor proteins and sequencing tags are ligated to the molecules. This library preparation step is fast — often under an hour with rapid kits.
The motor protein (typically a helicase) acts as a precise ratchet. It unwinds double-stranded regions if needed and controls the speed at which the single-stranded molecule threads through the nanopore — roughly 400–500 bases per second in current chemistries. This controlled pace ensures the electrical signal remains readable.
Voltage pulls the negatively charged molecule from the cis to the trans side of the membrane. As it enters the nanopore, the molecule begins its journey through the narrow channel.
Each passing k-mer alters the ionic current in a characteristic way, producing the raw data trace.
Real-time algorithms convert the electrical signals into a sequence. Raw data is stored as FAST5 files (containing the squiggle trace), which are then processed into standard FASTQ files containing the A, C, G, and T sequence.
At Quintara Bio, this entire process is streamlined so you simply submit your plasmid or amplicon samples and receive high-quality long-read data — often the next day.
The raw output of a nanopore sequencer is not a clean string of bases but a continuous graph of current intensity over time — affectionately known in the field as a "squiggle." This plot reflects the unique electrical fingerprint of overlapping k-mers (typically 5–9 bases) rather than individual nucleotides.
Because the pore reads several bases at once, the signal is a composite pattern that the basecalling software must deconvolute. The squiggle is rich with information, including subtle shifts caused by base modifications, which is why nanopore sequencing excels at direct epigenetic detection.
Basecalling is the critical computational step that transforms raw electrical data into usable sequence. Modern Oxford Nanopore basecallers, such as Dorado, use deep learning models trained on vast datasets of known sequences and their corresponding squiggles. These AI models run locally on the sequencing device or in the cloud and can process data in real time.
The output begins as FAST5 files that store the full raw signal. After basecalling, the software generates FASTQ files with quality scores. The latest models achieve impressive consensus accuracy, making nanopore data suitable for even demanding applications like clinical research or synthetic biology.
One of nanopore sequencing's most powerful features is its ability to detect epigenetic modifications directly. Because it sequences native molecules without PCR amplification, chemical changes such as methylation alter the way the base interacts with the pore and therefore change the electrical signal. These subtle differences are captured in the squiggle and can be called alongside the primary sequence.
This native sequencing capability opens doors to true multi-omic insights in a single run — something short-read technologies typically require separate, more complex protocols to achieve.
Unlike Illumina's sequencing-by-synthesis approach, which uses fluorescent labels and optical imaging, nanopore sequencing relies entirely on electrical signals. It also differs from Pacific Biosciences (PacBio) single-molecule real-time sequencing, which uses optical detection of fluorescent nucleotides in zero-mode waveguides.
The electrical detection method gives nanopore its unique combination of portability, real-time output, and direct modification detection — advantages that have made it the platform of choice for field work, rapid diagnostics, and long-read applications.
The electrical principle delivers several practical benefits:
· Real-time sequencing with data available within minutes
· Ultra-long reads that simplify assembly of complex genomes
· Highly portable devices suitable for labs or remote locations
· PCR-free workflows that preserve native modifications
· Direct RNA sequencing without conversion to cDNA
These features make nanopore sequencing especially valuable for plasmid verification, amplicon analysis, metagenomics, and full-length transcript studies.
No technology is perfect. Early nanopore systems had higher error rates, particularly in homopolymer regions where multiple identical bases in a row can produce similar signals. Signal noise and the complexity of interpreting overlapping k-mers also require robust computational resources.
However, these limitations have been steadily addressed through chemistry and software improvements, and many projects now achieve high consensus accuracy through moderate coverage.
Accuracy has advanced rapidly. The latest R10.4.1 pores combined with super-accurate (SUP) basecalling models routinely deliver Q20+ single-molecule accuracy and Q30+ consensus with duplex reads. Newer AI models continue to reduce errors in challenging sequence contexts, while higher-throughput flow cells increase yield without sacrificing read length.
In essence, the process is a beautiful chain of events:
DNA/RNA → motor protein control → nanopore translocation → ionic current disruption (squiggle) → AI basecalling → FASTQ sequence.
This direct electrical readout, combined with long reads and real-time output, explains why nanopore sequencing has become indispensable for modern genomics.
It measures unique disruptions in ionic current caused by each base (or k-mer) as the molecule passes through the nanopore. AI algorithms then decode these signals.
A steady flow of charged ions through the open nanopore. Bases temporarily block or alter this flow, creating the measurable electrical signal.
Yes — modern chemistry and basecalling routinely achieve >99% single-read accuracy and >99.9% consensus accuracy, with ongoing improvements.
The raw data format that stores the complete electrical squiggle trace before basecalling converts it to FASTQ.
At Quintara Bio, we make third-generation nanopore sequencing simple, fast, and affordable. Simply submit your plasmid DNA or amplicon samples, and our team will handle library preparation, sequencing on the latest Oxford Nanopore platforms, and data analysis — delivering high-quality long-read results often the next day.
Choose from our specialized services:
a. l Whole Plasmid Sequencing (WPS Express, WPS Standard, and WPS Pro) — perfect for routine verification, high-throughput construct screening, or resolving complex plasmid mixtures.
b. l Amplicon Sequencing (AmpValue, AmpStandard, and AmpPro) — ideal for amplicons covering different length, read depth, and mixed library verification.
AmpValue is our most cost-effective entry-level nanopore amplicon sequencing service, uniquely positioned in the market as a rapid “no-friction” solution for PCR validation. Unlike traditional workflows that often require purified DNA and additional preparation steps, AmpValue supports direct use of unpurified PCR products, significantly reducing experimental turnaround time and operational complexity.
This makes it one of the few nanopore-based amplicon sequencing services specifically optimized for high-throughput screening rather than only deep variant characterization. By combining low cost, overnight delivery, and minimal sample preparation, AmpValue fills a unique gap between Sanger sequencing and high-depth nanopore amplicon analysis — enabling researchers to quickly confirm experimental success before scaling into more advanced sequencing workflows.
Current Launch Promotions:
a. l Get 5 Free Samples on AmpValue Nanopore Sequencing (promo code: AmpValue5Free)
b. l Amplicon Sequencing Clean Up at only $1 per sample (promo code: Cleanup1)
Results are ready for download the next day, with same-day service available in the Cambridge/Boston area. Whether you need rapid, cost-effective long-read data or a custom project, our streamlined workflow and expert support ensure reliable, publication-ready insights without the hassle of owning hardware.
Order now through the QuinGo online portal or contact us for help with a custom order. Let Quintara Bio turn your raw samples into actionable genomic data with speed and confidence.